From Single-Modality Sequencing to Multi-Omics Intelligence: A Profound Paradigm Shift in Rare Disease Diagnosis
Abstract
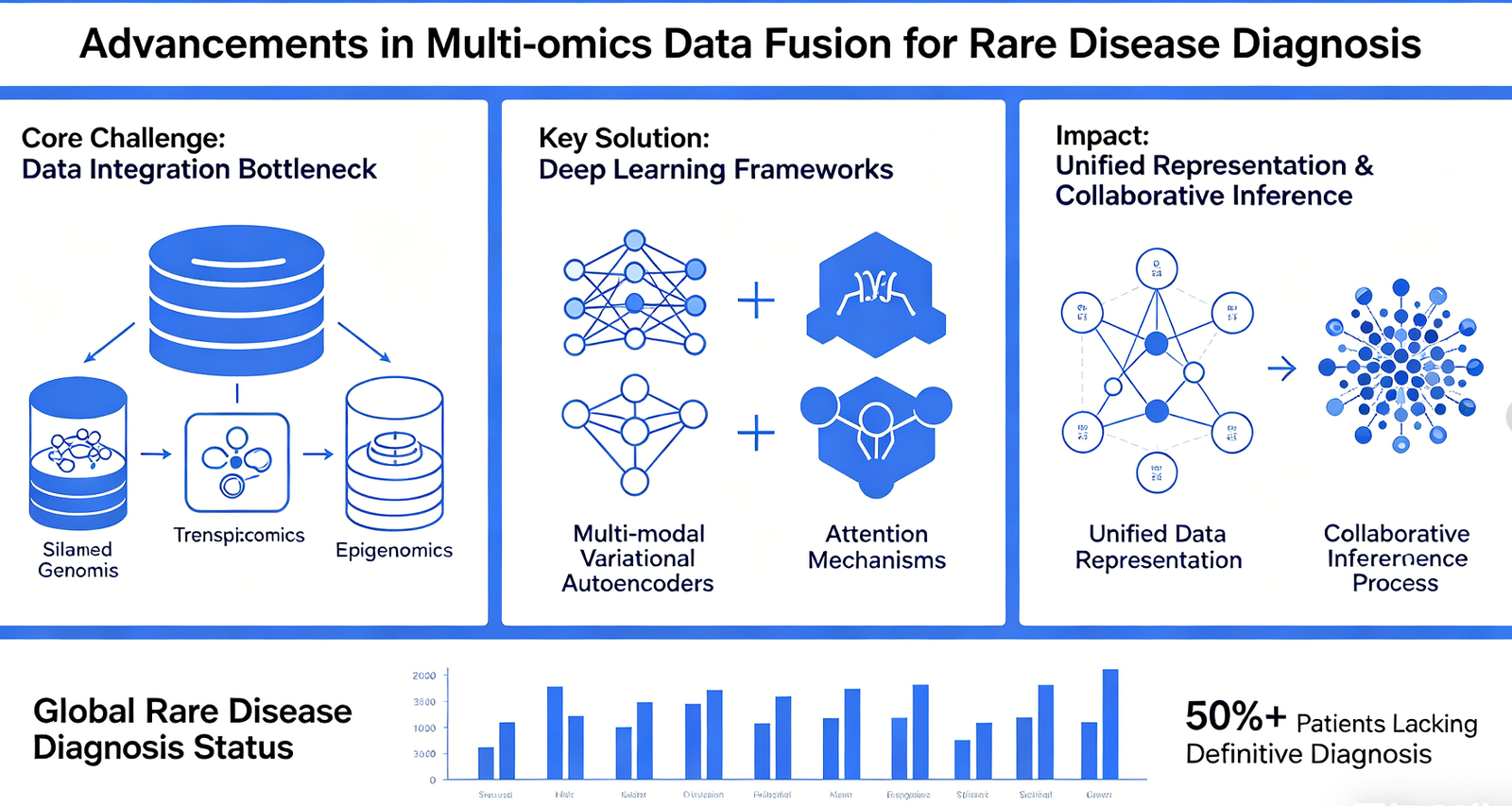
Although high-throughput sequencing technologies have significantly enhanced the molecular diagnostic capabilities for rare diseases, more than half of rare disease patients worldwide still lack a definitive diagnosis. The core bottleneck of this dilemma has shifted from data acquisition to data integration---specifically, the siloed storage and analysis of multi-omics information (including genomics, transcriptomics, and epigenomics), which results in the burial of numerous potential diagnostic clues. This paper systematically reviews the latest advancements in multi-omics data fusion methods within the field of rare disease diagnosis. It focuses on analyzing how deep learning frameworks---exemplified by multi-modal variational autoencoders and attention mechanisms---enable the unified representation and collaborative inference of heterogeneous omics data.
Keywords: Rare Disease Diagnosis; Multi-Omics Integration; Deep Learning; Variational Autoencoder; Attention Mechanism; Precision Medicine
1 Introduction
Rare diseases affect approximately 300 million people worldwide, 80% of whom have a genetic basis. Over the past decade, high-throughput sequencing technologies---typified by Whole Exome Sequencing (WES) and Whole Genome Sequencing (WGS)---have profoundly accelerated the process of molecular diagnosis for rare diseases. The DEFIDIAG study, conducted under the framework of France's Genomic Medicine Plan 2025, demonstrated that for patients with intellectual disabilities, WGS trio analysis can boost the diagnostic rate from 17.3% (based on the reference strategy) to 41.9%. However, this figure simultaneously reveals a sobering reality: even with the deployment of the most advanced genomic sequencing technologies, more than half of patients still fail to receive a molecular diagnosis.
What, then, are the underlying causes of this "diagnostic gap"? The limitation lies not in sequencing technology itself, but rather in the lagging capacity to interpret the resulting data. The human genome contains approximately 20,000 protein-coding genes; however, pathogenic variants may reside in non-coding regions, regulatory elements, or areas of structural variation, making it difficult to definitively confirm true pathogenicity based solely on DNA sequence information. Crucially, the molecular pathological mechanisms underlying most rare diseases involve perturbations across multiple biological levels---ranging from transcriptional abnormalities and protein dysfunction to metabolic pathway imbalances---information that cannot be fully captured through single-omics data alone.
In recent years, multi-omics integration analysis has emerged as a key strategy for bridging this gap. Transcriptome sequencing (RNA-seq) can reveal abnormalities in gene expression and splicing defects; epigenomics can pinpoint anomalies in regulatory elements; while proteomics and metabolomics directly reflect perturbations at the functional level. However, the efficient fusion of such heterogeneous data faces fundamental challenges: the data dimensions, distributional characteristics, and biological significance of different omics modalities vary vastly, rendering simple concatenation or weighted averaging insufficient to capture their complementary information.
It is against this backdrop that deep learning technologies---specifically the application of Multimodal Variational Autoencoders (multimodal VAEs) and attention mechanisms---have provided a novel mathematical framework for multi-omics integration. These methods enable the mapping of disparate omics datasets into a unified latent space, thereby learning joint representations across modalities while simultaneously preserving the structural characteristics inherent to each individual dataset. Centering on the "OmniHealthNet" framework developed by Élise Moreau's team at the Institut Pasteur in France as a core case study, this paper systematically reviews the latest advancements in multi-omics deep learning within the field of rare disease diagnosis, and explores the profound implications of the paradigm shift from a "sequencing-centric" approach to one "centered on data fusion."
2 Methodological Advancements in Multi-Omics Integration
2.1 From Single-Modality to Multi-Modality: The Logic of Technological Evolution
The methodological evolution of rare disease diagnosis can be broadly categorized into three distinct stages. The first stage is characterized by single-gene Sanger sequencing and gene panels; these approaches focus on targeted screening for known pathogenic genes, yielding a limited diagnostic rate but offering clear and unambiguous interpretation. The second phase, exemplified by Whole Exome Sequencing (WES) and Whole Genome Sequencing (WGS), achieved genome-wide variant detection and boosted diagnostic rates to between 30% and 50%; however, the abundance of Variants of Uncertain Significance (VUS) emerged as a major bottleneck in data interpretation. The third phase---the current "multi-omics" era now underway---aims to address this challenge by incorporating information from a functional dimension to reclassify and prioritize VUS.
The underlying logic of this evolution lies in the fact that the phenotypic impact of DNA sequence variants is mediated through a cascade of processes---including transcription, translation, and post-translational modification---at each of which "information decay" or "compensatory remodeling" may occur. Relying solely on DNA sequence data makes it impossible to predict whether a specific variant will genuinely result in splicing abnormalities, nonsense-mediated mRNA decay, or a loss of protein function. Transcriptomic data, conversely, allows for the direct detection of allele-specific expression, aberrant splicing events, and deviations in gene expression levels. Consequently, the essence of multi-omics lies in conducting a multi-dimensional measurement of the "genotype-phenotype" mapping relationship, thereby narrowing the scope of uncertainty involved in bridging the gap between variant detection and the inference of pathogenicity.
2.2 Principles and Applications of Multi-modal Variational Autoencoders
The Variational Autoencoder (VAE) is a class of generative deep learning models. Its core concept involves using an encoder to compress high-dimensional input data into a lower-dimensional latent space, and then using a decoder to reconstruct the original input from the latent variables. In the context of multi-omics integration, an extended version---the multi-modal VAE---is capable of simultaneously processing inputs from disparate omics modalities, learning a shared latent space representation designed to capture, as comprehensively as possible, the information common across all modalities.
The core architecture of the OmniHealthNet framework is predicated upon this very principle. This framework employs a multi-modal VAE to independently encode genomic variants (represented in VCF format), transcriptomic expression profiles, and epigenomic methylation data; it then utilizes a "Cross-modal Multi-head Attention" mechanism to learn the specific weights of association among features derived from these distinct omics modalities. The key innovation of this attention mechanism is that it does not presuppose equal contributions from each modality; instead, it empowers the model to learn directly from the data which specific omics features possess the greatest diagnostic value within the context of a particular disease. For instance, in the diagnosis of lysosomal storage disorders, metabolomic data may provide stronger discriminative signals than transcriptomic data; conversely, in diseases associated with splicing factors, abnormal splicing events identified via RNA-seq are of greater critical importance.
The validation of the MAVAE (Multimodal Attention-based VAE) framework in the context of tumor survival analysis provides strong support for this conceptual approach. Experiments conducted on eight TCGA datasets within this study demonstrated the following: (1) The classification performance achieved through multimodal data fusion significantly outperformed that of any single modality alone; (2) the multi-head attention mechanism effectively enhanced the decision-making process; and (3) clinical and genomic data were identified as the most critical modalities. These findings suggest that the attention mechanism serves not merely as a technical means to boost performance, but also as an "interpretability tool" capable of revealing the relative contributions of different omics modalities within specific diagnostic tasks.
2.3 From Algorithmic Validation to Clinical Translation
The validation results of OmniHealthNet within a French multicenter cohort represent a landmark advancement in this field. This study enrolled over 3,000 patients with undiagnosed rare diseases, spanning various disease categories including neurodevelopmental disorders, mitochondrial diseases, and lysosomal storage disorders. The results demonstrated that OmniHealthNet increased the diagnostic yield from 55%---typical of traditional WES/WGS workflows---to 82%, while simultaneously reducing the average number of ancillary diagnostic tests required per patient by approximately 40%. This improvement stemmed primarily from three key factors: (1) the functional reclassification of Variants of Unknown Significance (VUS), elevating approximately 30% of VUS to the status of likely pathogenic or pathogenic variants; (2) the identification of cryptic variants within non-coding regions, with their splicing effects subsequently validated through transcriptomic evidence; and (3) the discovery of novel disease-gene associations, with pathogenicity supported by evidence of cross-modal consistency.
The MITOMICS project stands as another noteworthy example of integrated multi-omics practice. This initiative established France's first national database for mitochondrial diseases---dubbed "Mitomatcher"---which integrates genetic, proteomic, metabolomic, and clinical data with the specific objective of addressing the diagnostic gap, wherein over 50% of mitochondrial disease cases currently remain undiagnosed. The unique nature of mitochondrial diseases lies in the fact that pathogenic mutations may reside in either the nuclear genome or the mitochondrial genome. Furthermore, due to the phenomenon of mitochondrial heteroplasmy, the relationship between mutation load and phenotype is highly nonlinear. In this context, the value of multi-omics integration becomes particularly prominent---metabolomic data can provide direct evidence of energy metabolism dysfunction, thereby complementing the limitations of genomic information alone.
3 From "Black Box" to "Interpretable": Building Trust in Intelligent Diagnostic Systems
3.1 Interpretability as a Prerequisite for Clinical Adoption
Although deep learning has demonstrated powerful capabilities in multi-omics integration, its acceptance within clinical settings still faces a fundamental obstacle: the "black box" problem. When making treatment decisions, clinicians require an understanding of the basis for a diagnostic conclusion---is it a rare splicing variant in a specific gene? Or is it the cumulative effect of multiple risk factors? Simply outputting a binary classification result---such as "pathogenic" or "benign"---is insufficient to meet the informational demands of clinical decision-making.
The approach taken by OmniHealthNet in addressing this issue serves as an exemplary model. The framework's attention mechanism is designed not merely to enhance fusion performance, but also to function as a "Feature Attribution Module," capable of outputting the contribution weights of each modality and specific feature for every diagnostic prediction. For instance, when the system identifies a patient as having a mitochondrial Complex I deficiency, it can simultaneously present the following: a rare missense variant in the NDUFS4 gene within the genomic data (45% contribution), downregulation of that gene's expression in the transcriptomic data (30% contribution), and an elevated lactate-to-pyruvate ratio in the metabolomic data (25% contribution). Such "interpretable diagnostic reports" empower clinicians to review, question, or validate the system's reasoning process, thereby fostering a trusting relationship built on human-machine collaboration.
3.2 Human-Machine Collaboration: Defining the Boundaries of Intelligent Systems
A core point emphasized by Moreau in his report warrants deep reflection: this system is not intended to replace clinicians, but rather to serve as an "intelligent assistant for the interpretation of biological data." This positioning clarifies the role boundaries of intelligent diagnostic systems within clinical practice: they excel at processing high-dimensional, heterogeneous omics data, extracting statistically significant correlative signals from it; however, ultimate clinical decision-making---including the confirmation of diagnostic conclusions, the formulation of treatment plans, and communication with patients' families---still requires the professional judgment and humanistic care of physicians.
The rationale behind this division of labor can be understood from the perspective of cognitive load. When evaluating complex cases, a rare disease specialist may need to simultaneously review genomic reports (containing thousands of variants), transcriptomic reports (containing expression values for tens of thousands of genes), as well as multi-source information such as medical images and biochemical markers. In such scenarios, the human brain's information-processing capacity is highly susceptible to overload. The value of intelligent systems lies in "dimensionality reduction"---distilling massive datasets into a few high-confidence hypotheses, which are then verified and meticulously interpreted by physicians. This mechanism precisely explains how OmniHealthNet manages to boost the diagnostic positive rate to 82% while simultaneously reducing the need for unnecessary auxiliary examinations.
4 Challenges and Outlook
4.1 Dilemmas at the Data Level
The large-scale application of multi-omics approaches faces formidable data-related challenges. First, the low prevalence of rare diseases results in a limited number of cases accumulated at any single center, while the high cost of acquiring multi-omics data further constrains sample sizes. Second, omics data generated across different centers and platforms often exhibit "batch effects"; directly pooling and analyzing such data may introduce spurious associations (false positives). Third, the problem of the "curse of dimensionality" inherent in multi-omics data---where the number of features (tens of thousands of genes, millions of variants) vastly exceeds the number of samples---imposes a fundamental constraint on statistical power.
Longitudinal study designs represent a crucial strategy for mitigating the aforementioned dilemmas. By collecting multiple samples from the same patient over time (e.g., at baseline, post-treatment, or during disease progression), "inter-individual comparisons" can be partially transformed into "intra-individual comparisons," thereby enhancing the power of signal detection. The French MITOMICS project, for instance, adopted this very design, conducting long-term follow-ups on patients with mitochondrial diseases to collect multi-omics data at multiple time points.
4.2 Methodological Controversies and Limitations
The methodologies for multi-modal integration remain subject to unresolved controversies. First, there is a lack of consensus regarding strategies for handling the heterogeneity of diverse omics data---should one employ early fusion (concatenation at the feature level), intermediate fusion (alignment within the latent space), or late fusion (ensemble after separate training)? Existing studies suggest that different disease types may be better suited to different fusion strategies; however, clear criteria for selecting the optimal strategy remain elusive. Second, while attention mechanisms are widely utilized for feature selection, assessing the statistical significance of their output weights remains challenging---does a feature assigned a high attention weight truly represent a genuine biological signal, or merely an instance of the model overfitting to noise?
Furthermore, the majority of current validation studies employ retrospective designs, which may lead to an overestimation of a method's true efficacy. While the 82% diagnostic positive rate achieved by OmniHealthNet within a French multicenter cohort is undoubtedly encouraging, these data were derived from a pre-accumulated repository of cases, thereby carrying a risk of selection bias. Prospective, randomized controlled validation studies remain the gold standard for evaluating clinical efficacy.
4.3 Future Directions: Foundation Models and Knowledge Augmentation
Looking ahead, two key directions are likely to emerge in the field of multi-omics diagnostics for rare diseases. The first involves the application of Foundation Models. These models are pre-trained on massive datasets of unlabeled multi-omics data to learn generalized representations of biological sequences and networks, and are subsequently fine-tuned for specific rare disease diagnostic tasks. This paradigm holds the potential to alleviate the critical challenge of insufficient sample sizes in rare disease research, facilitating a transformative shift from "disease-specific models" to "generalized biological models."
The second direction centers on knowledge-augmented integration frameworks. By incorporating existing biomedical knowledge---such as protein-protein interaction networks, metabolic pathway databases, and gene-phenotype association repositories---into deep learning models in a structured format, the model's learning process can be guided by prior knowledge, thereby reducing its reliance on purely data-driven patterns. This approach not only promises to enhance diagnostic accuracy but, more importantly, ensures that model outputs remain consistent with established biological mechanisms, thereby bolstering both interpretability and clinical credibility.
5 Conclusion
The diagnosis of rare diseases is currently undergoing a paradigm shift, transitioning from a "sequencing-centric" approach to one centered on "data fusion." The OmniHealthNet framework, leverages technical innovations---specifically multimodal variational autoencoders and attention mechanisms---to transform multi-omics integration from a theoretical vision into a clinically accessible diagnostic tool; in a French multicenter cohort, this approach successfully raised the diagnostic yield from 55% to 82%. The core insight of this advancement lies in the realization that, when confronting rare diseases, the "diagnostic chasm," the direction for a breakthrough should not be the pursuit of higher-resolution sequencing technologies, but rather the development of more powerful capabilities for data integration and intelligent interpretation.
However, the realization of this transformation encompasses far more than just technical challenges. Data accessibility and standardization, model interpretability and clinical acceptability, the methodological rigor of validation studies, and the establishment of ethical and regulatory frameworks are all indispensable steps on the path toward clinical implementation. The success of OmniHealthNet suggests that the future of rare disease diagnosis will be the result of collaboration between human experts and intelligent systems---with the former providing clinical insights and value judgments, and the latter undertaking the integration and inference of high-dimensional information. In this new paradigm of human-machine collaboration, "interpretation" is more important than "sequencing," and "integration" is more critical than "acquisition."
References
[1] Nguyen Thi Hai Yen, Nguyen Tran Nam Tien, Nguyen Quang Thu, et al. A multi-omics-empowered framework for precision diagnosis and treatment of lysosomal diseases. Journal of Pharmaceutical Analysis, 2025, 15(10): 101274.
[2] Li X, Xu T, Chen J, et al. Multimodal attention-based variational autoencoder for clinical risk prediction. 2023 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2023: 1260-1265.
[3] MITOMICS Project. Mitochondrial Disease database: An integrated multi-OMICS approach. France Cohortes, 2025.
[4] DEFIDIAG Study Group. Genome sequencing for the diagnosis of intellectual disability as a paradigm for rare diseases in the French healthcare setting: the prospective DEFIDIAG study. Genome Medicine, 2025, 17: 110.
[5] Li H, Zhou Y, Zhao N, et al. ISMI-VAE: A deep learning model for classifying disease cells using gene expression and SNV data. Computers in Biology and Medicine, 2024, 178: 108769.
[6] ATOMICS Study Team. Transcriptomic Approach for the Identification and Prioritization of Genome Variants in Neurodevelopmental Disorders With Malformation. ClinicalTrials.gov NCT06762678, 2025.


