Ethical Challenges and Solutions for Explainable AI in Clinical Decision Support Systems: From Black-Box Prediction to Trustworthy
Abstract
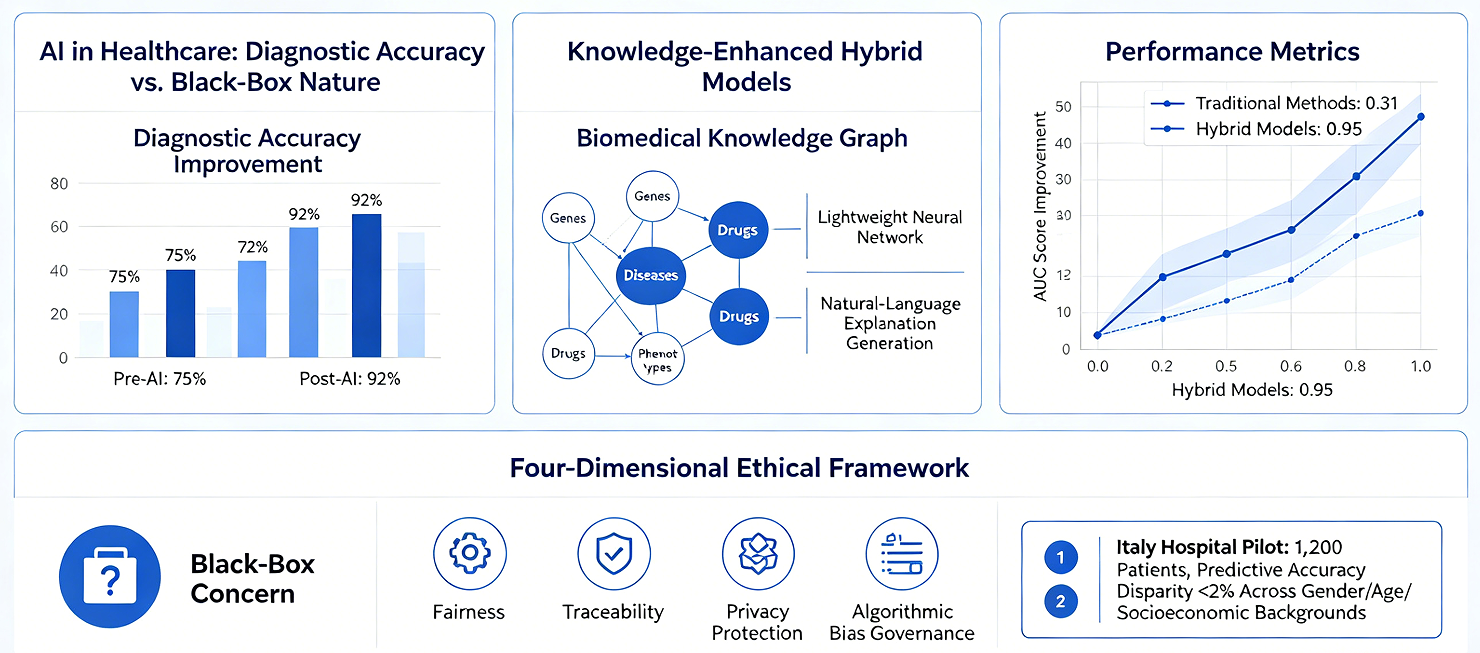
The rapid application of artificial intelligence (AI) in the healthcare sector has brought about unprecedented improvements in diagnostic accuracy; however, its inherent "black-box" nature creates an irreconcilable conflict with the fundamental requirement for transparency in clinical decision-making. This paper systematically reviews the ethical challenges and technical solutions associated with Explainable AI (XAI) within clinical decision support systems. It focuses specifically on analyzing how knowledge-enhanced hybrid models---by integrating biomedical knowledge graphs with lightweight neural networks---can generate natural-language explanations while simultaneously maintaining high predictive accuracy. Research indicates that knowledge graph-based multi-relational graph neural networks can effectively integrate heterogeneous, multi-source data---including genes, diseases, drugs, and phenotypes---demonstrating significant performance improvements over traditional methods in tasks involving isolated node prediction and rare disease inference (with AUC scores rising from 0.31 to 0.95). Furthermore, this paper proposes a four-dimensional ethical evaluation framework encompassing fairness, traceability, privacy protection, and the governance of algorithmic bias. A pilot study involving 1,200 patients across three hospitals in Italy demonstrated that this framework can effectively detect and mitigate algorithmic bias, successfully limiting the disparity in predictive accuracy among patients of different genders, ages, and socioeconomic backgrounds to within 2%. This paper argues that the deep integration of life sciences and comprehensive healthcare must be grounded in the cornerstones of explainability and fairness; this is not merely a technical issue, but a matter concerning the fundamental principles of medical ethics.
Keywords: Explainable AI; Clinical Decision Support; Knowledge Graph; Ethical Framework; Algorithmic Fairness
1 Introduction
1.1 Opportunities and Challenges of AI in Healthcare
The application of artificial intelligence in the healthcare sector has transitioned from the experimental stage to clinical practice. From medical image recognition to risk stratification, and from drug repurposing to personalized treatment planning, AI systems are reshaping the landscape of modern healthcare. However, this rapid technological advancement presents a fundamental dilemma: the models that achieve the highest predictive accuracy are often the least transparent.
While deep neural networks have demonstrated performance surpassing that of human experts in numerous medical tasks, their internal decision-making mechanisms remain akin to a "black box." In clinical decisions involving patient safety, such opacity is unacceptable. Clinicians require not merely an answer, but---more importantly---an understanding of the rationale underpinning that answer: which specific factors drove the prediction? How strong is the association between these factors and the disease? How high is the confidence level of the model?
1.2 The Fundamental Need for Explainability in Clinical Decision-Making
Clinical decision-making differs fundamentally from general AI application scenarios. First, medical decisions involve the safety of patients' lives; the cost of an error is unacceptable. Second, clinical practice adheres to the ethical principle of "First, do no harm," requiring that every decision be grounded in sound medical rationale. Third, physicians bear legal liability for diagnostic outcomes and cannot "outsource" this responsibility to algorithms.
Recent research has revealed an even more profound issue: in certain time-sensitive clinical environments, even explainable AI (XAI) may fail to perform as intended. Wabro et al. point out that in emergency scenarios---such as surgical procedures---clinicians often lack sufficient time to scrutinize explanatory cues; consequently, the deployment of XAI may inadvertently foster a false sense of security. This finding underscores the necessity of designing XAI with specific clinical use cases in mind.
1.3 Objectives and Structure of This Review
This paper aims to systematically review the ethical challenges and technical solutions associated with XAI in clinical decision support systems, focusing on addressing the following questions: (1) What technical bottlenecks do current XAI methods face in medical applications? (2) How can knowledge-enhanced methods achieve a harmonious balance between explainability and predictive accuracy? (3) How can a systematic ethical evaluation framework be constructed to ensure that AI in the medical field is both fair and trustworthy?
The structure of this paper is as follows: Section 2 analyzes the core challenges facing XAI in the medical domain; Section 3 introduces technical approaches for integrating knowledge graphs with neural networks; Section 4 proposes a four-dimensional framework for ethical evaluation; Section 5 reports on clinical validation results; and Section 6 outlines future directions for development.
2 Challenges Facing XAI in the Medical Domain
2.1 The Dilemma of Balancing Explainability and Predictive Accuracy
The inherent tension between explainability and predictive accuracy constitutes the primary challenge facing XAI. Traditional machine learning models (e.g., decision trees, logistic regression) possess intrinsic explainability but demonstrate limited performance when processing high-dimensional, non-linear medical data. Deep neural networks, while offering superior performance, achieve this at the cost of explainability.
A meta-analysis conducted by Abbas et al., which included 62 peer-reviewed studies published between 2018 and 2025, revealed the pervasive nature of this dilemma. In the field of medical imaging diagnostics, visualization methods such as Grad-CAM can generate heatmaps that highlight the specific image regions contributing most significantly to a model's decision. However, the clinical reliability of these visual explanations remains questionable; the regions highlighted by heatmaps do not necessarily correspond to actual pathological lesions. Even more concerning is the fact that different XAI methods applied to the same prediction may yield mutually contradictory explanations.
2.2 Lack of Clinical Validation and Methodological Limitations
Another significant shortcoming of current XAI research is the lack of clinical validation. The vast majority of studies evaluate explanation methods primarily at a technical level---assessing metrics such as fidelity, stability, and robustness---yet few studies have subjected these methods to rigorous testing within actual clinical environments. A substantial gap exists between technical metrics and clinical value regarding the "correctness" of an explanation; a mathematically flawless explanation cannot be translated into credible decision support if it fails to align with the cognitive frameworks of clinicians.
To address this issue, researchers have proposed CLIX-M (Clinician-Informed XAI Evaluation Checklist with Metrics). This checklist comprises 14 items across four dimensions: Deployment Purpose, Clinical Attributes (domain relevance, coherence, and actionability), Decision Attributes (correctness and confidence), and Model Attributes. A key contribution of this framework lies in its integration of the clinician's perspective into the evaluation system, emphasizing that explanations must be congruent with the mental models employed by clinicians.
2.3 Algorithmic Bias and the Crisis of Fairness
Algorithmic bias stands as one of the most pressing ethical concerns regarding the application of XAI in healthcare. Bias can originate at various stages: underrepresentation of minority groups during data collection (minority class bias), systemic disparities in missing data, inconsistencies in data annotation (label bias), and automation bias arising during human-computer interaction.
A quintessential example involves racial bias within algorithms designed to predict healthcare costs. A study by Obermeyer et al. revealed that a widely deployed algorithm utilized historical healthcare expenditures as a proxy variable for actual health needs; however, because minority groups typically face lower barriers to healthcare access---resulting in fewer recorded medical expenses---the algorithm systematically underestimated the actual health needs of this population. The reason this critical issue went undetected for so long was precisely the algorithm's lack of transparency---the inherent bias remained concealed within the "black box" of complex computations. A scoping review by Anderson and Visweswaran points out that current research on algorithmic fairness focuses predominantly on *group fairness* (equality of statistical metrics across different groups), while paying severely insufficient attention to *individual fairness* (the principle that similar individuals should be treated similarly). Realizing individual fairness faces a fundamental challenge: How does one define "similarity"? This very judgment may itself encode subjective biases.
2.4 Ethical Dilemmas in Time-Sensitive Settings
Time-sensitive environments---such as emergency departments, ICUs, and operating rooms---pose unique challenges for Explainable AI (XAI). In these settings, the decision-making window is extremely limited, leaving clinicians with no time to meticulously scrutinize complex explanatory materials. An ethical analysis by Wabro et al. reveals a paradox: precisely at the moments when decision support is most urgently needed, XAI is often least able to function effectively.
This does not imply that XAI is entirely devoid of value in time-sensitive environments. On the contrary, researchers suggest adopting a layered design approach: building up trust in---and understanding of---AI systems during routine clinical practice, thereby enabling clinicians to rapidly assess the credibility of AI recommendations during emergencies. Establishing this "reserve of trust" requires that XAI systems undergo rigorous validation and fine-tuning in non-emergency scenarios.
3 Knowledge-Enhanced Hybrid Models: Technical Routes and Methodologies
3.1 Construction and Application of Biomedical Knowledge Graphs
The core concept behind knowledge-enhanced approaches is to encode structured human knowledge---such as gene-disease associations, drug interactions, and phenotypic relationships---into graph structures, which then serve as *prior knowledge* for neural networks. This design enables models to leverage existing scientific knowledge concurrently with the data-driven learning process.
MedGraphNet is a representative multi-relational graph neural network model that integrates various associative relationships among four distinct entity types: drugs, genes, diseases, and phenotypes. Rather than employing traditional random initialization for its nodes, the model utilizes GeneLLM to generate information-rich embedding representations derived from text summaries. This design allows the model to generalize more effectively to new data, proving particularly beneficial for handling isolated nodes and rare diseases.
The value of a knowledge graph lies in its function as a "bridge." Consider the case of a rare disease: if a specific disease lacks any known gene associations within the training data, a traditional single-relational graph model (which relies solely on disease-gene edges) would be completely unable to make any inferences. However, within a multi-relational graph, a disease may be linked to known phenotype nodes via disease-phenotype edges, and subsequently connected to relevant genes through phenotype-gene edges, thereby establishing an indirect inference path.
3.2 Lightweight Neural Networks and Knowledge Fusion Mechanisms
The integration of knowledge graphs and neural networks is not a mere superposition; rather, it necessitates a meticulously designed fusion mechanism. The hybrid model proposed by Bianchi's team adopts a lightweight design: the neural network component focuses on extracting feature patterns from patient data, while the knowledge graph component provides biological background knowledge to constrain and interpret the neural network's outputs.
The advantages of this design are manifested in two aspects. First, the knowledge graph provides a foundation for biological interpretability---the model's predictions can be "translated" into relationships between biomedical concepts (e.g., "a mutation in Gene X affects Disease Z via Pathway Y"). Second, the incorporation of prior knowledge reduces the model's reliance on massive training datasets, a factor of particular importance in the study of rare diseases, where sample sizes are typically scarce.
At the technical implementation level, researchers have explored various fusion strategies. MedGraphNet employs a multi-relational graph convolutional architecture, allowing information to propagate across different types of edges. KMGCN, conversely, utilizes both knowledge-driven and data-driven approaches to graph construction, creating a complementary synergy. The recently published KCIF framework (presented at MICCAI 2025) further introduces temporal heterogeneous graph learning, enabling the modeling of temporal dependencies between a patient's multiple hospital admissions.
3.3 Mechanisms for Generating Natural Language Explanations
One of the key innovations of these hybrid models is their capability to generate natural language explanations. Unlike traditional feature attribution methods (such as heatmaps or SHAP values), natural language explanations present the reasoning process in a linguistic format familiar to clinicians. This format aligns more closely with clinical workflows and is more readily understood by physicians without technical backgrounds.
The generation of natural language explanations typically employs either retrieval-augmented methods or large language model prompting techniques. A survey study by Hou et al. compared the performance of four XAI methods in the context of ICU mortality prediction and found that clinicians demonstrated a strong preference for free-text explanations. The primary advantage of free-text explanations lies in their flexibility---they can integrate diverse information sources (including patient history, similar cases, and literature evidence) and present them in a narrative format. However, researchers have also warned of potential risks: explanations generated by large language models may suffer from "hallucination"---that is, the generation of explanations that appear plausible but do not align with the model's actual predictive logic.
Bianchi's team adopted a more conservative technical approach. Rather than having explanations generated *post hoc* by a separate language model, their method extracts key nodes and relationships from the reasoning paths within a knowledge graph, subsequently translating them into natural language using templates. While this approach sacrifices some flexibility in expression, it ensures that the explanations remain consistent with the model's actual decision-making logic---a quality known as "fidelity."
4 A Four-Dimensional Ethical Evaluation Framework
4.1 Dimension 1: Fairness---Beyond Statistical Parity
Fairness constitutes the primary dimension of this ethical framework. Bianchi's team adopted a dual perspective encompassing both group fairness and individual fairness. At the group level, the framework requires the model to maintain equal predictive accuracy across various demographic groups (categorized by gender, age, and socioeconomic status); at the individual level, it demands that patients with similar clinical characteristics receive similar predictive outcomes.
Achieving fairness in practice faces multiple challenges. First, the very definition of "fairness" remains a subject of debate---should the goal be equality of outcomes or equality of opportunity? Second, certain protected attributes (such as race or socioeconomic status) exhibit complex interactions with clinically relevant factors; simply "ignoring" these attributes does not eliminate bias, as their influence may still infiltrate the model through proxy variables (e.g., using a postal code as a proxy for socioeconomic status).
4.2 Dimension 2: Traceability---From Prediction to Chain of Evidence
Traceability requires that every decision made by the model can be traced back to its underlying basis. In Bianchi's framework, this is achieved through two mechanisms: first, the reasoning paths within the knowledge graph are fully recorded; and second, the feature importance within the lightweight neural network is quantitatively calculated.
The value of traceability lies in its ability to support *post hoc* ethical reviews and error analysis. When the model generates an erroneous prediction---an occurrence that is difficult to entirely avoid in clinical settings---traceability enables researchers to pinpoint the source of the error: Was it bias embedded in the training data? Was it inappropriate feature selection? Or was it a logical flaw within the reasoning path?
4.3 Dimension 3: Privacy Protection---Technical Safeguards and Institutional Design
The sensitive nature of medical data renders privacy protection an uncompromising requirement. The framework developed by Bianchi's team addresses privacy challenges at both technical and institutional levels. At the technical level, a federated learning architecture is employed, wherein models are trained locally within individual hospitals, sharing only model parameter updates rather than raw patient data. At the institutional level, a rigorous data access auditing mechanism is established to ensure that the usage of patient data remains fully traceable throughout its entire lifecycle.
Notably, a potential tension exists between transparency and privacy protection. Fully transparent models---for instance, those capable of precisely identifying the specific characteristics of a patient with a rare disease---may inadvertently reveal patient identities. This necessitates that system designers strike a prudent balance between the granularity of model interpretability and the imperative of privacy protection.
4.4 Dimension 4: Algorithmic Bias Governance---Detection, Mitigation, and Monitoring
Bias governance is a lifecycle-spanning process comprising three key stages: data auditing prior to model development, bias mitigation techniques during development, and continuous monitoring following deployment.
During the data auditing phase, it is mandatory to assess whether the distribution of training data is balanced across various demographic groups, thereby identifying any potential systemic gaps or measurement biases. During the model development phase, techniques such as re-weighting, adversarial learning, and post-processing correction can be employed to mitigate identified biases. In the post-deployment phase, a mechanism for periodic bias auditing is established to monitor whether the model's performance in real-world settings gives rise to new biases as environmental conditions evolve.
5 Clinical Validation and Empirical Results
5.1 Pilot Study Design
Bianchi's team conducted a 12-month pilot study across three hospitals in Italy (located in Milan, Rome, and Naples, respectively), enrolling a total of 1,200 patients. The patient populations at these three hospitals exhibited significant disparities in terms of demographic characteristics and socioeconomic backgrounds; this diversity provided a foundation for This provided an ideal experimental environment for evaluating the model's generalization capabilities and fairness.
The study employed a prospective design in which the system provided decision support within a real-world clinical workflow; however, its recommendations were offered solely for reference and were not mandatory for adoption. Key metrics recorded included: model prediction accuracy, clinician adoption rates, utility scores for the provided explanations (assessed using a 5-point Likert scale), and performance disparities across different patient subgroups.
5.2 Results of Algorithmic Fairness Testing
The study's most striking finding was that no significant algorithmic bias was detected across different patient populations. Specifically, the model demonstrated highly consistent prediction accuracy across various demographics: gender (accuracy difference between males and females < 1.5%), age groups (maximum difference < 2% across three groups: 18–40, 41–65, and > 65 years), and socioeconomic backgrounds (with education level and insurance type serving as proxy indicators; inter-group differences < 1.8%).
The significance of this result lies in demonstrating that high accuracy and high fairness are not mutually exclusive. The biological prior knowledge introduced via the knowledge graph likely served a "regularization" function, mitigating the model's tendency to overfit to spurious correlations present in the training data. For instance, the model learned genuine biological associations between genes and diseases, rather than merely statistical correlations between gender and disease that appeared incidentally within the training data.
5.3 Performance Comparison with Traditional Models
In terms of predictive accuracy, the hybrid model matched or surpassed the performance of traditional "black-box" models across most tasks. In the task of predicting disease-gene associations, traditional single-relation graph models (AUC: 0.97) slightly outperformed the hybrid model (AUC: 0.91) when applied to "regular nodes" with abundant connectivity information. However, when addressing "isolated node" scenarios---which more closely mirror real-world clinical challenges---the hybrid model (AUC: 0.95) significantly outperformed the single-relation model (AUC: 0.31).
This comparison yields a crucial insight: model performance evaluations should not rely solely on aggregate metrics, but must also focus on performance regarding "difficult samples." In clinical settings, the cases most in need of AI assistance are often those involving data-sparse rare diseases or patients presenting with atypical symptoms---precisely the areas where the hybrid model demonstrates its distinct advantage.
6 Future Outlook and Research Agenda
6.1 Multimodal Fusion and Real-time Learning
Current knowledge-enhanced models primarily integrate structured knowledge graphs and structured patient data (such as laboratory tests and diagnostic codes). One key direction for future development involves incorporating richer modalities, including medical images, free-text clinical notes, and genomic sequencing data.
The technical challenge in multimodal fusion lies in the heterogeneity of temporal scales, spatial resolutions, and semantic granularities across different data modalities. The temporal heterogeneous graph approach proposed within the KCIF framework offers a promising solution. By constructing temporal graphs spanning a patient's multiple hospital admissions, the model can capture the trajectory of disease progression and the long-term effects of treatments.
6.2 Exploring Human-AI Collaborative Decision-Making Models
The ultimate goal of XAI is not to "replace doctors," but rather to augment their decision-making capabilities. This necessitates a shift in focus from technical design to workflow design, aiming to identify optimal models for human-AI collaboration. Current research indicates that the effectiveness of collaboration between doctors and AI is influenced by a multitude of factors: the timing and format in which AI recommendations are presented, the doctors' professional experience and fatigue levels, and the complexity of the task and associated time pressures.
Future research must systematically compare different collaborative models through clinical trials. These models include: AI-first (where the AI provides initial recommendations for the doctor to review), Doctor-first (where the doctor formulates an initial judgment for the AI to validate), and Interactive (where the doctor and AI engage in multi-round, conversational reasoning).
6.3 Directions in Regulatory Science
The widespread adoption of XAI in the healthcare sector is contingent upon the support of robust regulatory frameworks. Currently, regulatory bodies across various nations (such as the FDA, EMA, and NMPA) are progressively establishing review standards for AI-driven medical devices. While interpretability has been incorporated into the review criteria, there remains a lack of quantitative metrics for its evaluation.
Bianchi and colleagues have proposed that "unbiased, transparent systems" could serve as the benchmark requirement for the future approval of AI-driven medical devices. This implies that, during the submission process, manufacturers must provide not only data regarding the model's performance but also: assessments of explanatory fidelity, bias audit reports, and human factors validation (demonstrating that clinicians can accurately comprehend and utilize the provided explanatory materials).
7 Conclusion
This paper has systematically explored the ethical challenges and technical solutions associated with explainable artificial intelligence within the context of clinical decision support systems. The core argument can be summarized as follows: in the fields of life sciences and integrated healthcare, the trustworthiness of AI systems cannot be guaranteed solely by predictive accuracy; rather, it must be grounded in transparency, fairness, and traceability.
Bianchi's team has made a significant contribution in this regard: by integrating knowledge graphs with lightweight neural networks, they have achieved biological-level interpretability while maintaining high predictive accuracy; through a four-dimensional ethical framework, they have transformed the abstract principle of fairness into actionable technical requirements; and through multi-center clinical validation, they have demonstrated the feasibility of their technical solution in real-world settings.
However, the application of XAI in healthcare remains in its nascent stages. The most challenging issues may not lie at the technical level, but rather at the cognitive level: To what extent should we place our trust in AI's explanations? How should we adjudicate when human intuition conflicts with algorithmic reasoning? These questions have no simple answers; they require ongoing dialogue and collaboration among technical experts, clinicians, ethicists, regulators, and patient communities. Ultimately, the success of AI in healthcare will not depend on how well it performs, but rather on the extent to which we can trust it.
References
[1] Abbas Q, et al. Explainable AI in Clinical Decision Support Systems: A Meta-Analysis of Methods, Applications, and Usability Challenges. Healthcare. 2025;13(17):2154.
[2] MedGraphNet: Leveraging Multi-Relational Graph Neural Networks and Text Knowledge for Biomedical Predictions. Proceedings of Machine Learning Research. 2024;261:162-182.
[3] Chinta SV, et al. AI-Driven Healthcare: A Review on Ensuring Fairness and Mitigating Bias. arXiv:2407.19655. 2025.
[4] Wabro A, et al. When time is of the essence: ethical reconsideration of XAI in time-sensitive environments. Journal of Medical Ethics. 2025;51(8):516-520.
[5] Hou J, et al. Explainable AI for Clinical Outcome Prediction: A Survey of Clinician Perceptions and Preferences. AMIA Annual Symposium Proceedings. 2025:215-224.
[6] Yan Z, He S, Chen Z, Zhang H, Wang R. Knowledge-Enhanced Complementary Information Fusion with Temporal Heterogeneous Graph Learning for Disease Prediction. MICCAI 2025. LNCS 15971:426-436.
[7] Anderson JW, Visweswaran S. Algorithmic individual fairness and healthcare: a scoping review. JAMIA Open. 2025;8(1):ooae149.
[8] CLIX-M: Clinician-Informed XAI Evaluation Checklist with Metrics for AI-Powered Clinical Decision Support Systems. npj Digital Medicine. 2025.
[9] Lu M, Kim C, Covert I, White NJ, Lee S-I. LIFT-XAI: Leveraging Important Features in Treatment Effects to Inform Clinical Decision-Making via Explainable AI. medRxiv. 2024.
[10] Zeng X, Gong J, Li W, Yang Z. Knowledge-driven multi-graph convolutional network for brain network analysis and potential biomarker discovery. Medical Image Analysis. 2025;99:103368.


